scRNAseq_combined_03_Batch_Clustering
retogerber
2024-02-12
Last updated: 2024-02-12
Checks: 7 0
Knit directory: synovialscrnaseq/
This reproducible R Markdown analysis was created with workflowr (version 1.6.2). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20210105) was run prior to running the code in the R Markdown file. Setting a seed ensures that any results that rely on randomness, e.g. subsampling or permutations, are reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version 58eeb06. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for the analysis have been committed to Git prior to generating the results (you can use wflow_publish or wflow_git_commit). workflowr only checks the R Markdown file, but you know if there are other scripts or data files that it depends on. Below is the status of the Git repository when the results were generated:
Ignored files:
Ignored: '/
Ignored: .Rhistory
Ignored: .Rproj.user/
Ignored: .empty/
Ignored: analysis/.Rhistory
Ignored: analysis/iSEE_interactive_document.html
Ignored: code/test_files/
Ignored: data/Culemann/
Ignored: data/E-MTAB-8322/
Ignored: data/Synovial scRNA-seq samples - Sheet1.csv
Ignored: data/Zhang_top20_singlecell_cluster_markers_fromGithub.csv
Ignored: data/findMarkers_results.rds
Ignored: data/findMarkers_results_v2.rds
Ignored: data/info/
Ignored: data/syn_sce_tidy_filtered.rds
Ignored: data/syn_sce_tidy_hvg.rds
Ignored: data/syn_sce_tidy_hvg_cms.rds
Ignored: docs/
Ignored: output/Figures_Paper/
Ignored: output/Sample_summaries_RA_comparisons.rds
Ignored: output/Sample_summaries_direct_dissociation.rds
Ignored: output/Sample_summaries_exvivo_treatment.rds
Ignored: output/Suppl_Figure_4d.rds
Ignored: output/barcodes.txt
Ignored: output/barcodes_filtered.txt
Ignored: output/column_metadata_filtered.txt
Ignored: output/combined_v7_SingleR_markers.rds
Ignored: output/combined_v7_SingleR_predictions.rds
Ignored: output/combined_v7_SingleR_predictions_lclc.rds
Ignored: output/combined_v7_SingleR_predictions_reclc.rds
Ignored: output/combined_v7_SingleR_predictions_recrec.rds
Ignored: output/combined_v7_SingleR_trained.rds
Ignored: output/combined_v7_sce.rds
Ignored: output/combined_v7_sce_filtered.rds
Ignored: output/combined_v7_sce_hvg.rds
Ignored: output/combined_v7_sce_hvg_cms.rds
Ignored: output/combined_v7_sce_hvg_cms_annotated.rds
Ignored: output/combined_v7_sce_tmp_hvg_cms.rds
Ignored: output/combined_v7_upsetplot_genelists.rds
Ignored: output/count_matrix_filtered.mtx
Ignored: output/count_matrix_unfiltered.mtx
Ignored: output/emptyDrops_result_v4.rds
Ignored: output/emptyDrops_result_v4_tmp.rds
Ignored: output/emptyDrops_result_v4tmptmp.rds
Ignored: output/findMarkers_results_v6.rds
Ignored: output/findMarkers_results_v6_ec.rds
Ignored: output/findMarkers_results_v6_main.rds
Ignored: output/findMarkers_results_v6_mp.rds
Ignored: output/findMarkers_results_v6_sf.rds
Ignored: output/findMarkers_results_v6_tc.rds
Ignored: output/findMarkers_results_v7_ec.rds
Ignored: output/findMarkers_results_v7_main.rds
Ignored: output/findMarkers_results_v7_mp.rds
Ignored: output/findMarkers_results_v7_sf.rds
Ignored: output/findMarkers_results_v7_tc.rds
Ignored: output/genes.txt
Ignored: output/genes_filtered.txt
Ignored: output/goana_results_v6_ec.rds
Ignored: output/goana_results_v6_mp.rds
Ignored: output/preprocessing_number_of_cells.rds
Ignored: output/syn_v4_sce_emptyDrops_invivo.rds
Ignored: output/syn_v4_swappedDrops_24300_after.rds
Ignored: output/syn_v4_swappedDrops_24300_before.rds
Ignored: output/syn_v4_swappedDrops_24793_after.rds
Ignored: output/syn_v4_swappedDrops_24793_before.rds
Ignored: output/syn_v6_cluster_cellid_match_invivo.rds
Ignored: output/syn_v6_clustering_lookup_invivo.rds
Ignored: output/syn_v6_clustering_lookup_multiple_invivo.rds
Ignored: output/syn_v6_sce.rds
Ignored: output/syn_v6_sce_Figure8.rds
Ignored: output/syn_v6_sce_Figure8_dic_ls.rds
Ignored: output/syn_v6_sce_ec_invivo.rds
Ignored: output/syn_v6_sce_filtered_invivo.rds
Ignored: output/syn_v6_sce_hdf5/
Ignored: output/syn_v6_sce_hvg_cms_doublet_invivo.rds
Ignored: output/syn_v6_sce_hvg_cms_doublet_subcluster_invivo.rds
Ignored: output/syn_v6_sce_hvg_invivo.rds
Ignored: output/syn_v6_sce_hvg_marker_genes.rds
Ignored: output/syn_v6_sce_mp_invivo.rds
Ignored: output/syn_v6_sce_sf_invivo.rds
Ignored: output/syn_v6_sce_tc_invivo.rds
Ignored: output/syn_v6_sfig1.rds
Ignored: output/syn_v6_vst_out_invivo.rds
Ignored: output/syn_v7_cluster_cellid_match_invivo.rds
Ignored: output/syn_v7_clustering_lookup_invivo.rds
Ignored: output/syn_v7_clustering_lookup_multiple_invivo.rds
Ignored: output/syn_v7_sce.rds
Ignored: output/syn_v7_sce_Figure8.rds
Ignored: output/syn_v7_sce_Figure8_dic_ls.rds
Ignored: output/syn_v7_sce_ec_invivo.rds
Ignored: output/syn_v7_sce_ec_invivo_trajectory.rds
Ignored: output/syn_v7_sce_ec_invivo_trajectory2.rds
Ignored: output/syn_v7_sce_ec_invivo_trajectory2_ATres.rds
Ignored: output/syn_v7_sce_ec_invivo_trajectory_icMat.rds
Ignored: output/syn_v7_sce_filtered_invivo.rds
Ignored: output/syn_v7_sce_hdf5/
Ignored: output/syn_v7_sce_hvg_cms_doublet_invivo.rds
Ignored: output/syn_v7_sce_hvg_cms_doublet_subcluster_invivo.rds
Ignored: output/syn_v7_sce_hvg_cms_doublet_subcluster_invivo_cleaned.rds
Ignored: output/syn_v7_sce_hvg_invivo.rds
Ignored: output/syn_v7_sce_mp_invivo.rds
Ignored: output/syn_v7_sce_sf_invivo.rds
Ignored: output/syn_v7_sce_tc_invivo.rds
Ignored: output/syn_v7_sfig1.rds
Ignored: output/syn_v7_vst_out_invivo.rds
Untracked files:
Untracked: analysis/clean_and_save_sce.R
Untracked: analysis/description_integration_wei_stephenson
Untracked: analysis/scRNAseq_complete_01_preprocessing_comparison.Rmd
Untracked: analysis/scRNAseq_complete_05_ec_trajectory_analysis.Rmd
Untracked: analysis/scRNAseq_complete_05_ec_trajectory_analysis_2.Rmd
Untracked: analysis/scRNAseq_complete_05_ec_trajectory_analysis_3.Rmd
Untracked: code/plot_utilities.Rmd
Untracked: code/rebuild_ezRun.R
Untracked: code/tmp1.R
Untracked: code/tmp1.Rmd
Untracked: nonhosted_public/
Untracked: singRstudio.sh.bak
Unstaged changes:
Modified: analysis/scRNAseq_combined_06_Figures.Rmd
Modified: analysis/scRNAseq_complete_04-2_celltype_markers.Rmd
Modified: analysis/scRNAseq_complete_04-2_celltype_markers_subcelltypes.Rmd
Modified: analysis/scRNAseq_complete_04_Annotation_v7.Rmd
Modified: analysis/scRNAseq_complete_Figures.Rmd
Modified: analysis/write_tsv.Rmd
Modified: code/create_hdf5.R
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the repository in which changes were made to the R Markdown (analysis/scRNAseq_combined_03_Batch.Rmd) and HTML (public/scRNAseq_combined_03_Batch.html) files. If you’ve configured a remote Git repository (see ?wflow_git_remote), click on the hyperlinks in the table below to view the files as they were in that past version.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| Rmd | 58eeb06 | Reto Gerber | 2023-05-30 | add new version |
| html | 58eeb06 | Reto Gerber | 2023-05-30 | add new version |
| Rmd | 4575ba6 | Reto Gerber | 2022-12-21 | Update analyis |
Set up
suppressPackageStartupMessages({
library(dplyr)
library(ggplot2)
library(purrr)
library(stringr)
library(SummarizedExperiment)
library(SingleCellExperiment)
library(scater)
library(scran)
library(igraph)
library(scuttle)
library(bluster)
library(BiocParallel)
})
n_workers <- 10
RhpcBLASctl::blas_set_num_threads(n_workers)
bpparam <- BiocParallel::MulticoreParam(workers=n_workers, RNGseed = 123)
analysis_version <- 7
here::here()[1] "/home/retger/Synovial/synovialscrnaseq"source(here::here("code","utilities_plots.R"))
set.seed(100)fig_ls <- list(sub=list(),all=list())load data from dim reduction
syn_sce_hvg_1 <- readRDS(file = here::here("output",paste0("combined_v",analysis_version,"_sce_hvg.rds")))
tmpfilename <- paste0("syn_v",analysis_version,"_sce_hvg_cms_doublet_subcluster_invivo.rds")
syn_sce_hvg_2 <- readRDS(file = here::here("output",tmpfilename))
t1 <- syn_sce_hvg_1
syn_sce_hvg_1 <- t1
t2 <- syn_sce_hvg_2
syn_sce_hvg_2 <- t2
# combine data
tmpind <- rowData(syn_sce_hvg_1)$ID %in% rowData(syn_sce_hvg_2)$IDLoading required package: tidySingleCellExperiment
Attaching package: 'tidySingleCellExperiment'The following object is masked from 'package:IRanges':
sliceThe following object is masked from 'package:S4Vectors':
renameThe following object is masked from 'package:matrixStats':
countThe following objects are masked from 'package:dplyr':
bind_cols, bind_rows, countThe following object is masked from 'package:stats':
filtertmpind2 <- rowData(syn_sce_hvg_2)$ID %in% rowData(syn_sce_hvg_1)$ID
syn_sce_hvg_1 <- syn_sce_hvg_1[tmpind,]
syn_sce_hvg_2 <- syn_sce_hvg_2[tmpind2,]
genematch <- match(rowData(syn_sce_hvg_2)$ID,rowData(syn_sce_hvg_1)$ID)
genematch <- genematch[!is.na(genematch)]
all(rowData(syn_sce_hvg_1)$ID[genematch] == rowData(syn_sce_hvg_2)$ID)[1] TRUEsyn_sce_hvg_1$main_celltype <- NA
syn_sce_hvg_1$minor_celltype <- NA
colData(syn_sce_hvg_2) <- colData(syn_sce_hvg_2)[,colnames(colData(syn_sce_hvg_2))[colnames(colData(syn_sce_hvg_2)) %in% colnames(colData(syn_sce_hvg_1))]]
colData(syn_sce_hvg_1) <- colData(syn_sce_hvg_1)[,colnames(colData(syn_sce_hvg_1))[colnames(colData(syn_sce_hvg_1)) %in% colnames(colData(syn_sce_hvg_2))]]
rowData(syn_sce_hvg_2) <- rowData(syn_sce_hvg_2)[,colnames(rowData(syn_sce_hvg_2))[colnames(rowData(syn_sce_hvg_2)) %in% colnames(rowData(syn_sce_hvg_1))]]
rowData(syn_sce_hvg_1)$is_hvg <- NULL
rowData(syn_sce_hvg_2)$is_hvg <- NULL
assay(syn_sce_hvg_2,"reconstructed") <- NULL
reducedDims(syn_sce_hvg_1) <- list()
reducedDims(syn_sce_hvg_2) <- list()
syn_sce <- cbind(syn_sce_hvg_2, syn_sce_hvg_1[genematch,])
colnames(syn_sce) <- make.unique(colnames(syn_sce),".")saveRDS(syn_sce, file = here::here("output",paste0("combined_v",analysis_version,"_sce_tmp_hvg_cms.rds")))combined HVG
all_gene_var <- modelGeneVar(syn_sce,
block=syn_sce$Sample)Warning in regularize.values(x, y, ties, missing(ties), na.rm = na.rm):
collapsing to unique 'x' values
Warning in regularize.values(x, y, ties, missing(ties), na.rm = na.rm):
collapsing to unique 'x' values
Warning in regularize.values(x, y, ties, missing(ties), na.rm = na.rm):
collapsing to unique 'x' values
Warning in regularize.values(x, y, ties, missing(ties), na.rm = na.rm):
collapsing to unique 'x' values
Warning in regularize.values(x, y, ties, missing(ties), na.rm = na.rm):
collapsing to unique 'x' valueshvg <- getTopHVGs(all_gene_var, fdr.threshold=0.05)
length(hvg)[1] 2643mean_var_comb <- map(unique(syn_sce$Sample), ~ {
all_gene_var$per.block[[.x]] %>%
as_tibble %>%
mutate(row_names = rownames(all_gene_var$per.block[[.x]]),
is_hvg = row_names %in% hvg,
Sample = .x)
}) %>%
purrr::reduce(rbind)
mean_var_comb %>%
ggplot() +
geom_point(aes(x = mean, y = total, color= is_hvg)) +
geom_line(aes(x=mean, y= tech)) +
labs(y="Variance",x="Mean expression") +
facet_wrap(~Sample)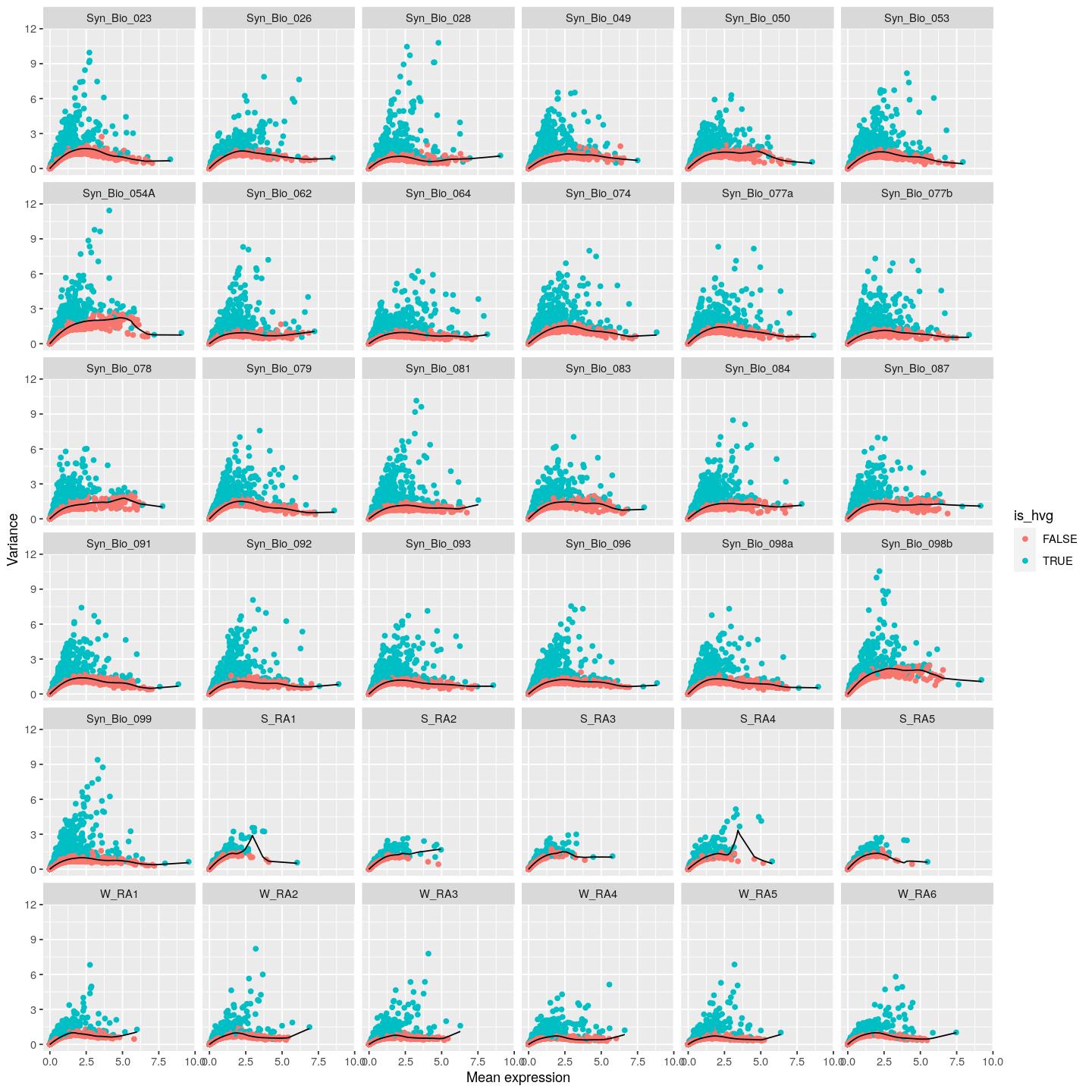
| Version | Author | Date |
|---|---|---|
| 58eeb06 | Reto Gerber | 2023-05-30 |
hvg_row_name <- "is_hvg"
rowData(syn_sce)[[hvg_row_name]] <- rownames(syn_sce) %in% hvgDimensionality reduction
use intrinsicDimension to get number of PC’s to keep. Run UMAP on reduced PCA.
set.seed(124)
sce_tmp <- syn_sce[rowData(syn_sce)[[hvg_row_name]],] %>%
runPCA(name = "PCA")
ndims <- intrinsicDimension::maxLikGlobalDimEst(as.matrix(reducedDim(sce_tmp, "PCA")), k=20)
reducedDim(sce_tmp,"PCA_reduced") <- reducedDim(sce_tmp,"PCA")[,seq_len(ceiling(ndims$dim.est))]
reducedDimNames(sce_tmp)[1] "PCA" "PCA_reduced"ncol(reducedDim(sce_tmp,"PCA_reduced"))[1] 17set.seed(100)
sce_tmp <- sce_tmp %>%
runUMAP(name = "UMAP", dimred = "PCA_reduced")
reducedDim(syn_sce, "PCA") <- reducedDim(sce_tmp,"PCA")
reducedDim(syn_sce, "PCA_reduced") <- reducedDim(sce_tmp,"PCA_reduced")
reducedDim(syn_sce, "UMAP") <- reducedDim(sce_tmp,"UMAP")Batch effects
Run batch correction using batchelor. Test batch correction using CellMixS.
syn_sce_cms <- syn_sce
bpstart(bpparam)
temp_sce <- batchelor::multiBatchNorm(
syn_sce,
batch = syn_sce$Sample,
subset.row = rownames(syn_sce)[rowData(syn_sce)[[hvg_row_name]]],
normalize.all = TRUE,
BPPARAM = bpparam
)
bpstop(bpparam)
merge_order <-
list(
list("S_RA5","S_RA2","S_RA4","S_RA1","S_RA3"), # stephenson samples
list(
list("Syn_Bio_053","Syn_Bio_054A",
"Syn_Bio_062","Syn_Bio_064"),#Peripheral_Spondyloarthritis
list("Syn_Bio_023","Syn_Bio_079","Syn_Bio_092"),#Psoriatic_Arthritis
list("Syn_Bio_083","Syn_Bio_084"),#Rheumatoid_Arthritis
list("Syn_Bio_074"),#Seronegative_Polyarthritis
list("Syn_Bio_026","Syn_Bio_028",
list("Syn_Bio_077b","Syn_Bio_077a"),
"Syn_Bio_049","Syn_Bio_050", "Syn_Bio_081",
"Syn_Bio_093","Syn_Bio_096",
list("Syn_Bio_098a","Syn_Bio_098b")
),#Seronegative_Rheumatoid_Arthritis
list("Syn_Bio_091","Syn_Bio_099"),#Spondyloarthritis
list("Syn_Bio_087"),#To_be_determined
list("Syn_Bio_078")#Undiff._Polyarthritis
), # own samples
list("W_RA1","W_RA2","W_RA3","W_RA4","W_RA5","W_RA6") # Wei samples
)
stopifnot(all(unlist(merge_order) %in% unique(syn_sce$Sample)))
stopifnot(all(unique(syn_sce$Sample) %in% unlist(merge_order)))
bpstart(bpparam)
temp_sce <- batchelor::fastMNN(temp_sce, batch=temp_sce$Sample, prop.k=0.05, merge.order = merge_order,
subset.row = rownames(temp_sce)[rowData(temp_sce)[[hvg_row_name]]], correct.all=TRUE,
BPPARAM = bpparam)
bpstop(bpparam)
assay(syn_sce_cms, "reconstructed") <- assay(temp_sce, "reconstructed")
reducedDim(syn_sce_cms, "corrected") <- reducedDim(temp_sce, "corrected")
bpstart(bpparam)
syn_sce_cms <- runUMAP(syn_sce_cms, dimred = "corrected", name = "UMAP_corrected",
BPPARAM = bpparam)
bpstop(bpparam)bpstart(bpparam)
syn_sce_cms <- CellMixS::cms(syn_sce_cms, k=100, group = "Sample",
dim_red = "corrected", res_name = "MNN",
BPPARAM = bpparam)
bpstop(bpparam)
CellMixS::visHist(syn_sce_cms)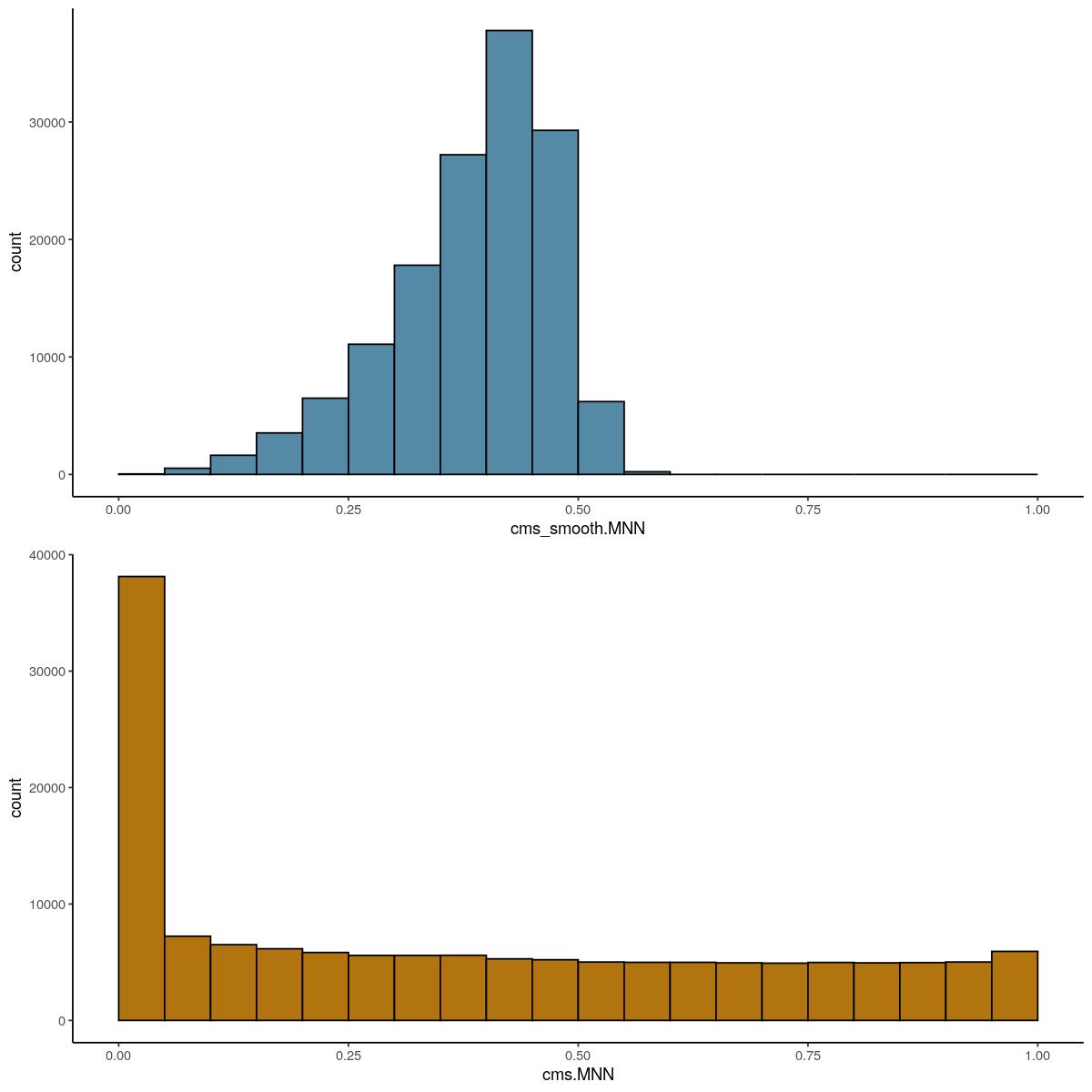
| Version | Author | Date |
|---|---|---|
| 58eeb06 | Reto Gerber | 2023-05-30 |
set.seed(123)
shuffle <- sample(seq_len(dim(syn_sce_cms)[2]))
cat("### PCAs Integration {.tabset}\n\n")PCAs Integration
cat("#### corrected \n\n")corrected
plotReducedDim(syn_sce_cms[,shuffle], "corrected", colour_by = "Sample")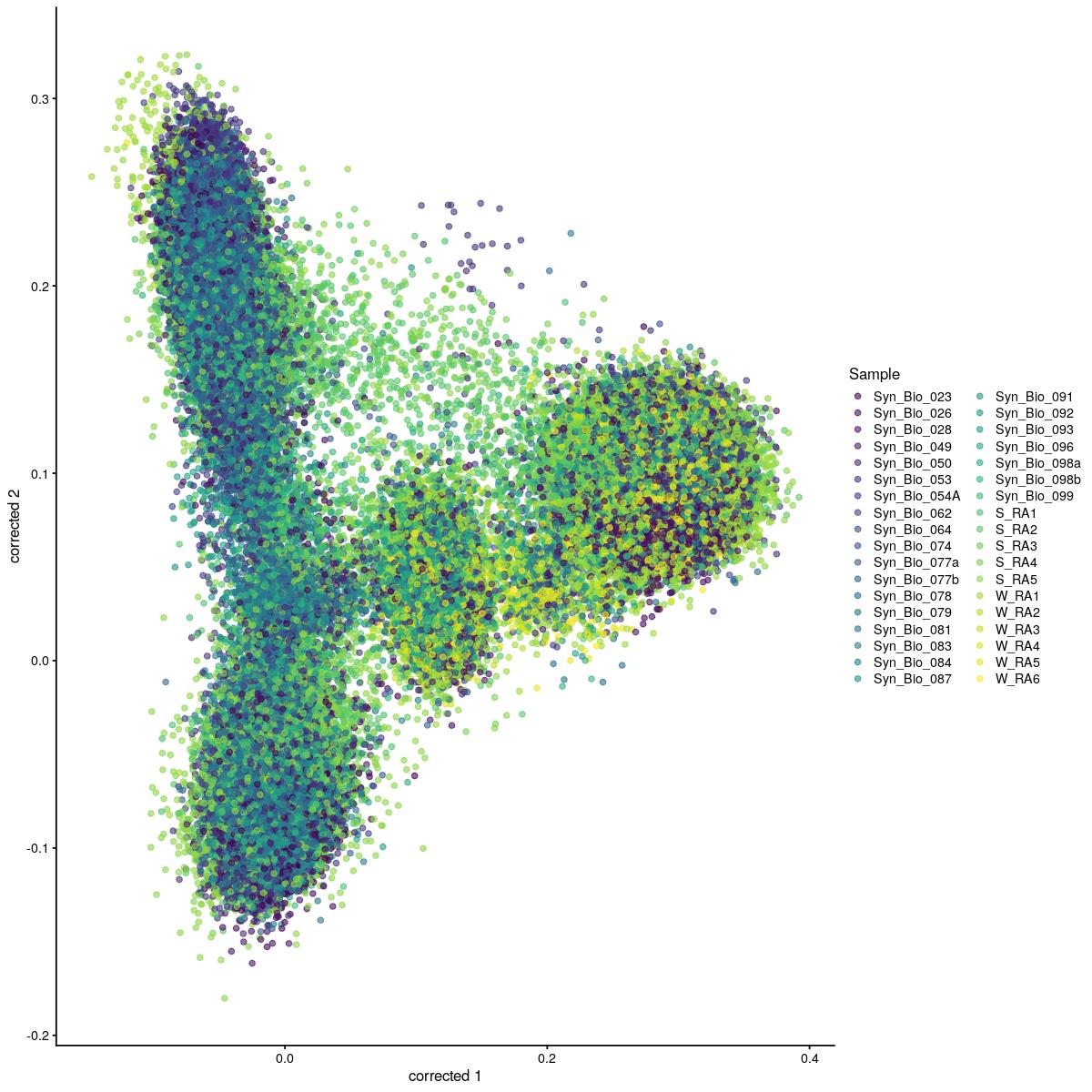
| Version | Author | Date |
|---|---|---|
| 58eeb06 | Reto Gerber | 2023-05-30 |
cat("#### uncorrected \n\n")uncorrected
plotReducedDim(syn_sce_cms[,shuffle], "PCA", colour_by = "Sample")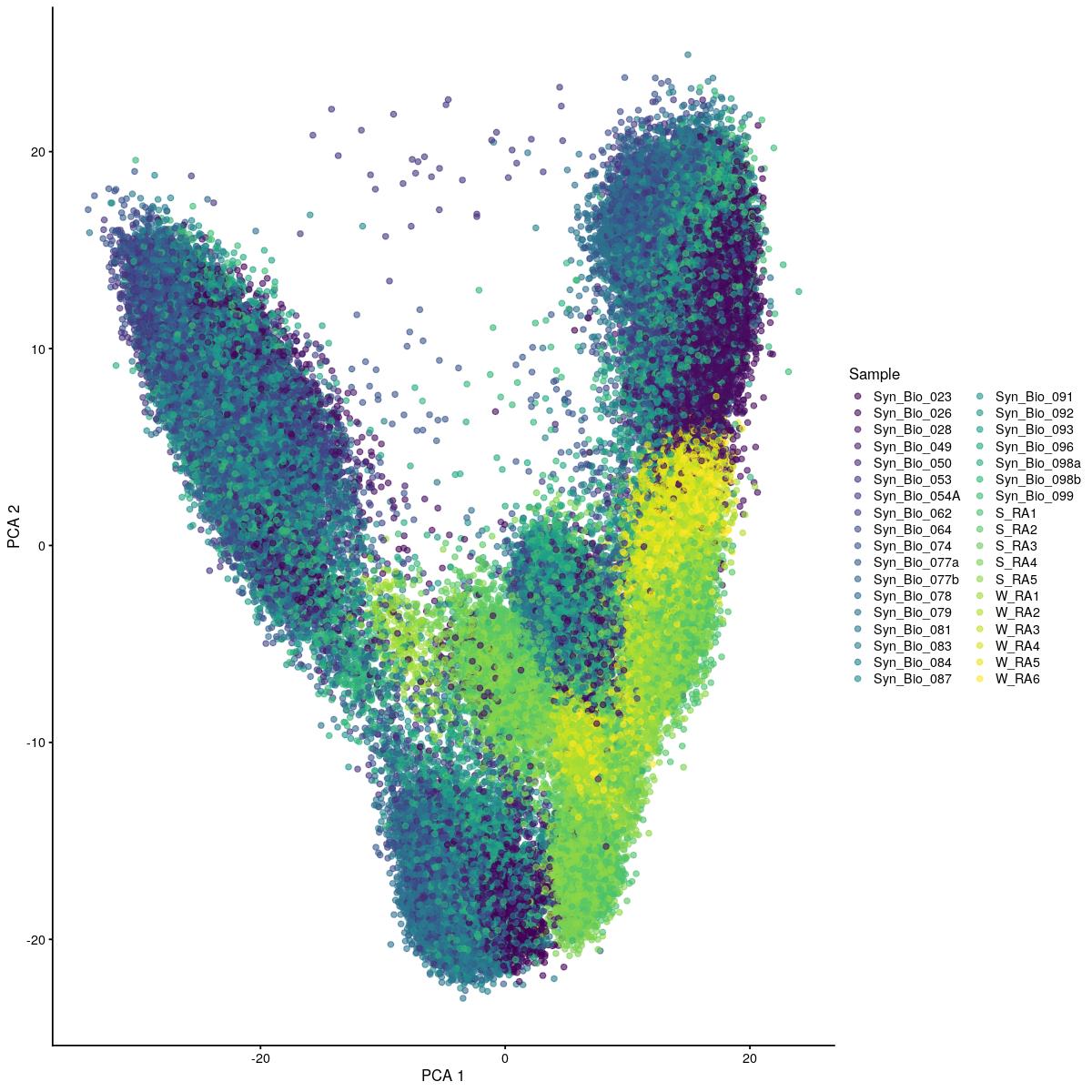
| Version | Author | Date |
|---|---|---|
| 58eeb06 | Reto Gerber | 2023-05-30 |
cat("#### corrected \n\n")corrected
plotReducedDim(syn_sce_cms[,shuffle], "corrected", colour_by = "Protocol")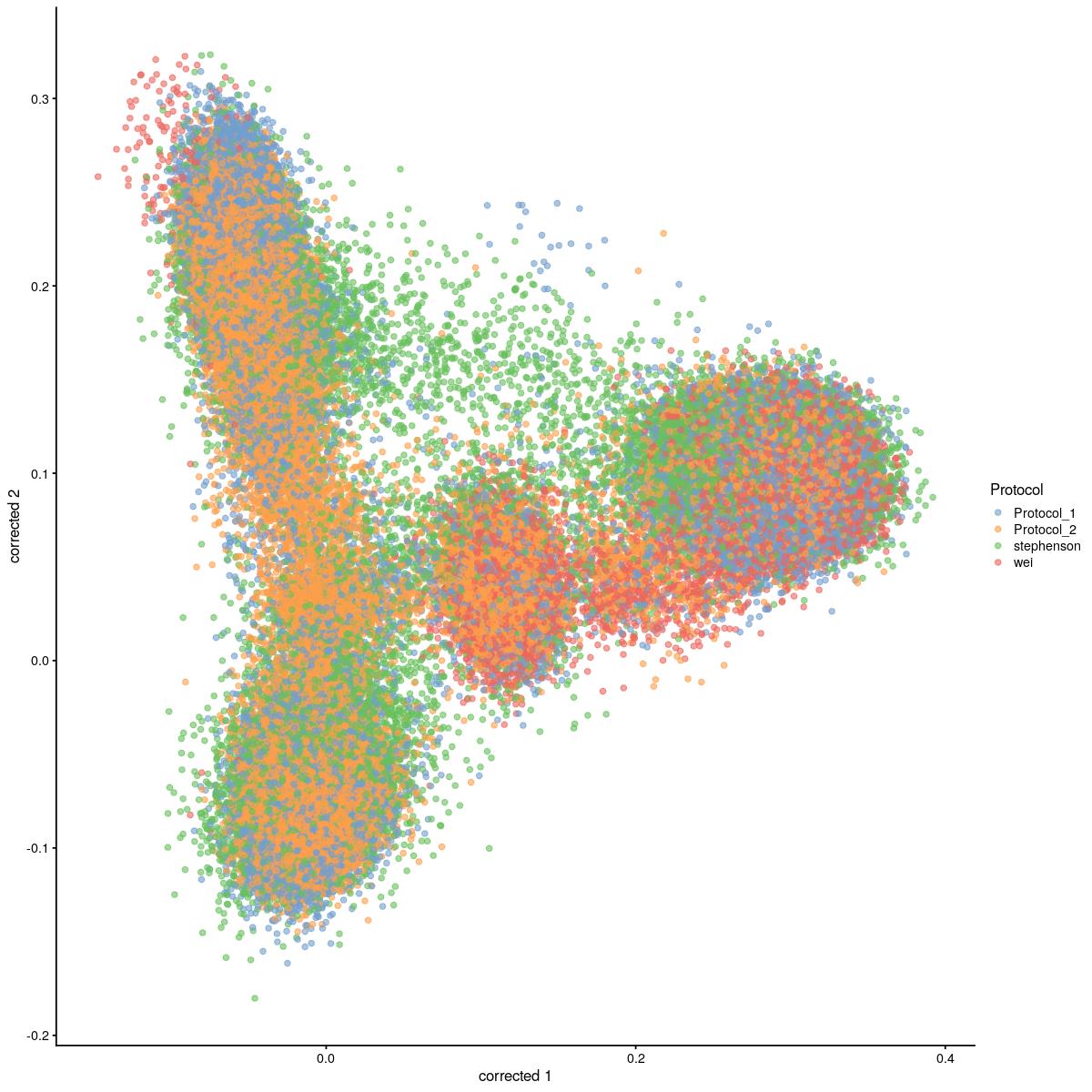
| Version | Author | Date |
|---|---|---|
| 58eeb06 | Reto Gerber | 2023-05-30 |
cat("#### uncorrected \n\n")uncorrected
plotReducedDim(syn_sce_cms[,shuffle], "PCA", colour_by = "Protocol")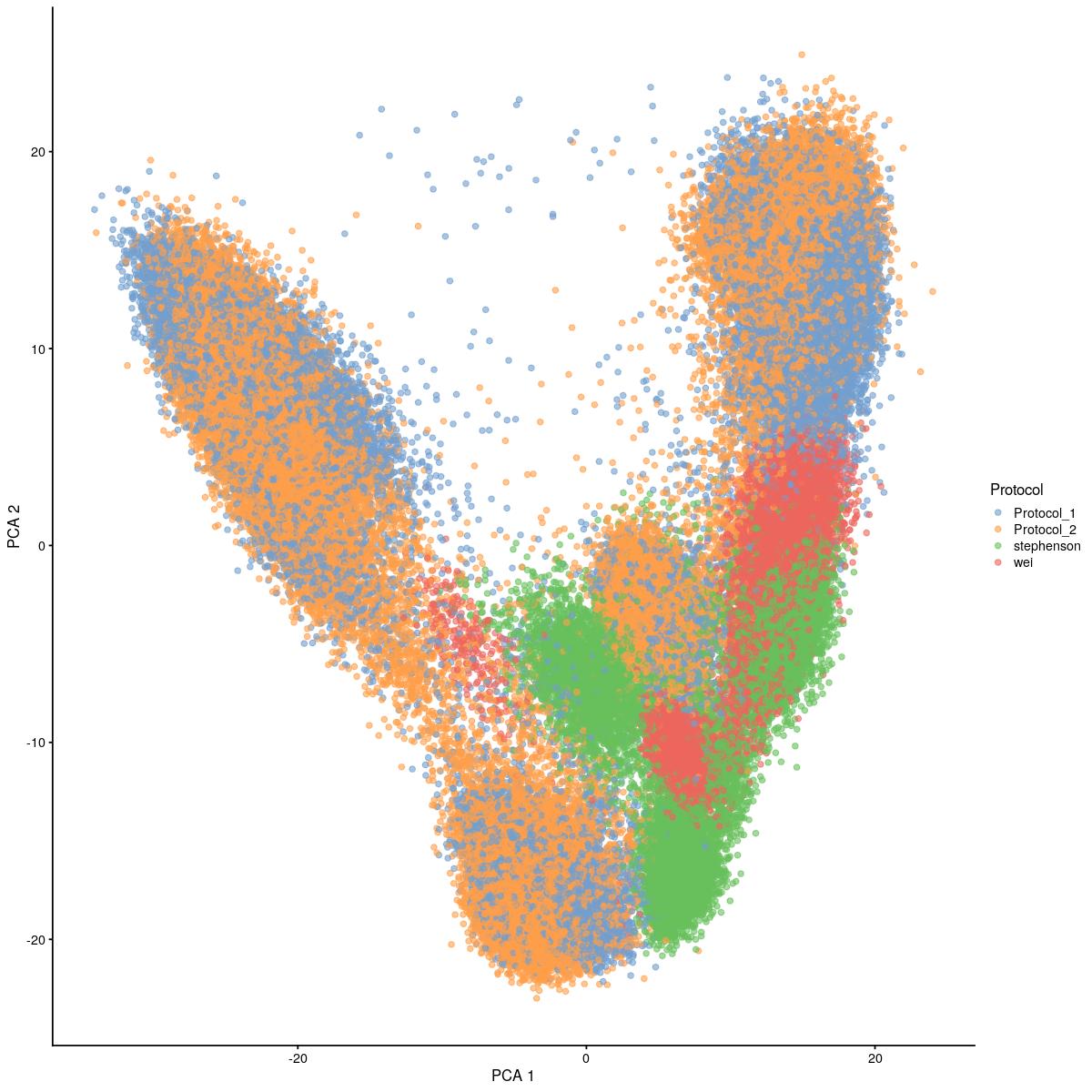
| Version | Author | Date |
|---|---|---|
| 58eeb06 | Reto Gerber | 2023-05-30 |
cat("\n\n### {-}")pltls <- purrr::map(unique(syn_sce_cms$Protocol),~{
tmpsce <- syn_sce_cms[,syn_sce_cms$Protocol==.x]
set.seed(123)
rsam <- sample(seq_along(tmpsce$Sample))
plotReducedDim(tmpsce[,rsam], "UMAP_corrected", colour_by = "Sample") +
labs(title=.x)
})
ggpubr::ggarrange(plotlist=pltls,common.legend = TRUE)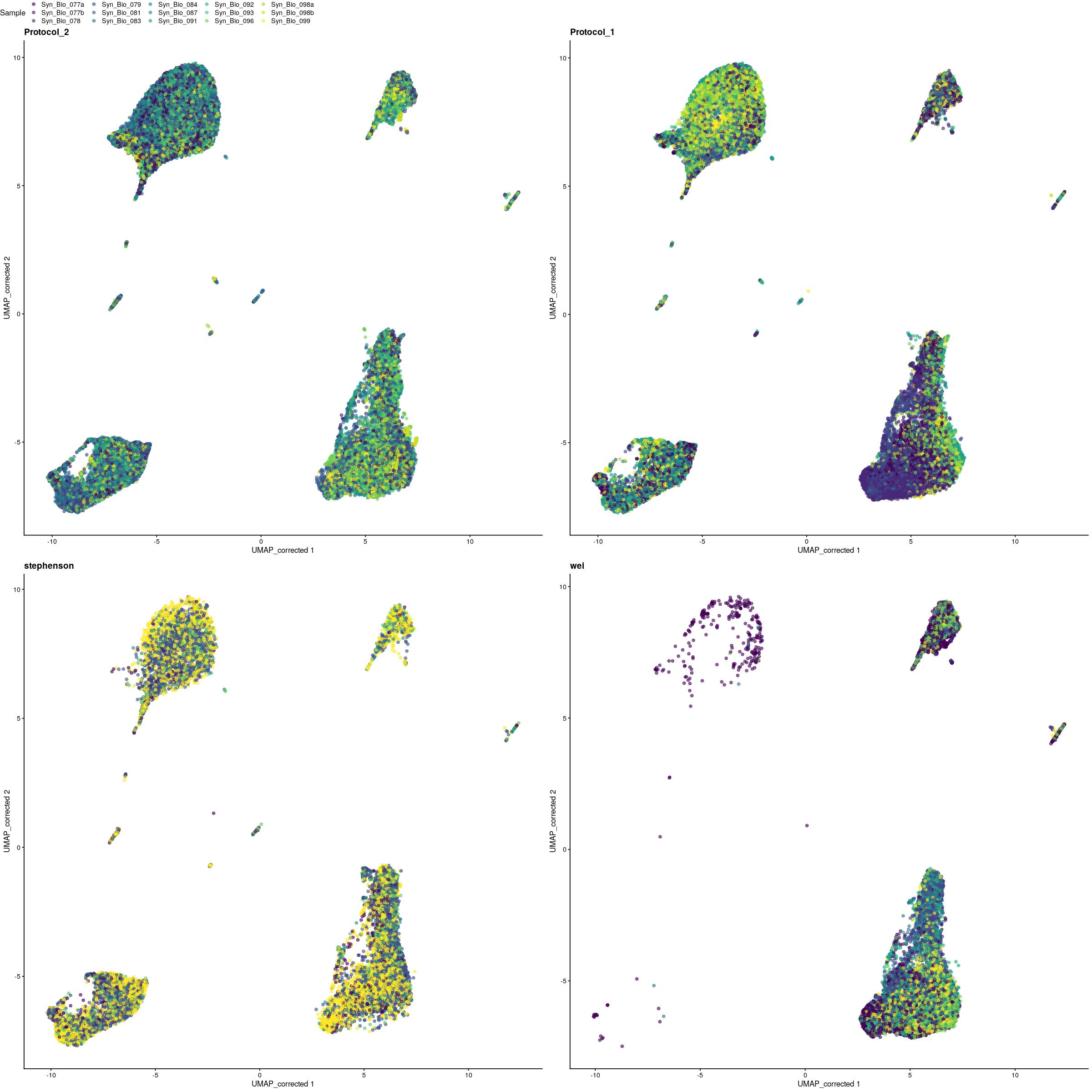
| Version | Author | Date |
|---|---|---|
| 58eeb06 | Reto Gerber | 2023-05-30 |
tmpsce_sub <- syn_sce_cms[,syn_sce_cms$Protocol=="stephenson"]
pltls <- purrr::map(unique(tmpsce_sub$Sample),~{
tmpsce <- tmpsce_sub[,tmpsce_sub$Sample==.x]
set.seed(123)
rsam <- sample(seq_along(tmpsce$Sample))
plotReducedDim(tmpsce[,rsam], "UMAP_corrected", colour_by = "Sample") +
labs(title=.x)
})
ggpubr::ggarrange(plotlist=pltls,common.legend = TRUE)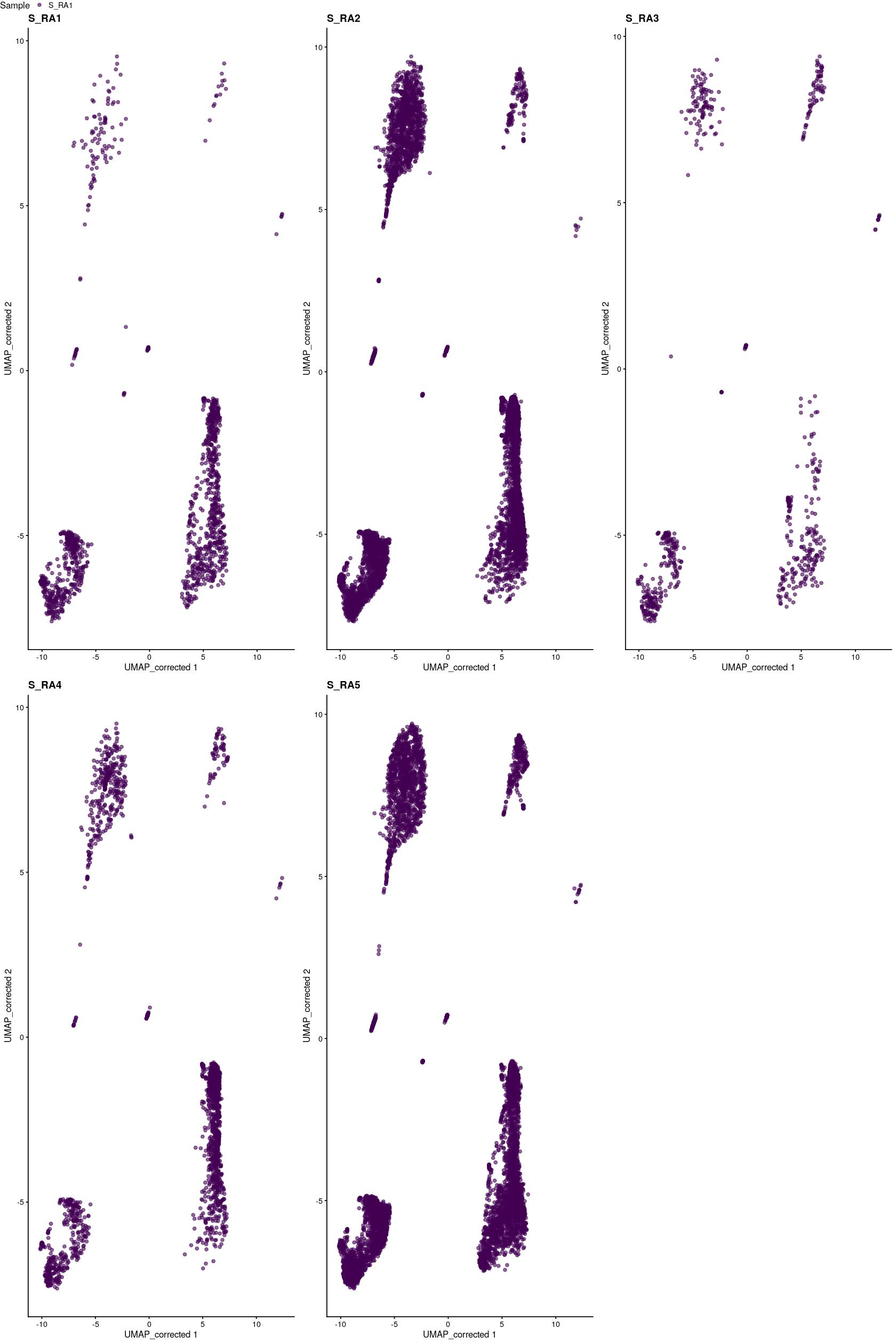
| Version | Author | Date |
|---|---|---|
| 58eeb06 | Reto Gerber | 2023-05-30 |
set.seed(123)
shuffle <- sample(seq_len(dim(syn_sce_cms)[2]))
fig_ls[['all']]$sfig2c1 <- plotReducedDim(syn_sce_cms[,shuffle], "UMAP_corrected", colour_by = "Sample") +
labs(color="Sample") +
scale_color_manual(values=sample_cols(unique(syn_sce_cms$Sample), n_split=5)) +
main_plot_theme()Scale for 'colour' is already present. Adding another scale for 'colour',
which will replace the existing scale.fig_ls[['all']]$sfig2c2 <- plotReducedDim(syn_sce_cms[,shuffle], "UMAP_corrected", colour_by = "Protocol") +
labs(color="Protocol") +
scale_color_manual(values=sample_cols(unique(syn_sce_cms$Protocol), n_split=2, palette=viridis::viridis)) +
main_plot_theme()Scale for 'colour' is already present. Adding another scale for 'colour',
which will replace the existing scale.cat("### UMAP corrected {.tabset}\n\n")UMAP corrected
cat("#### color by Sample \n\n")color by Sample
print(fig_ls[['all']]$sfig2c1)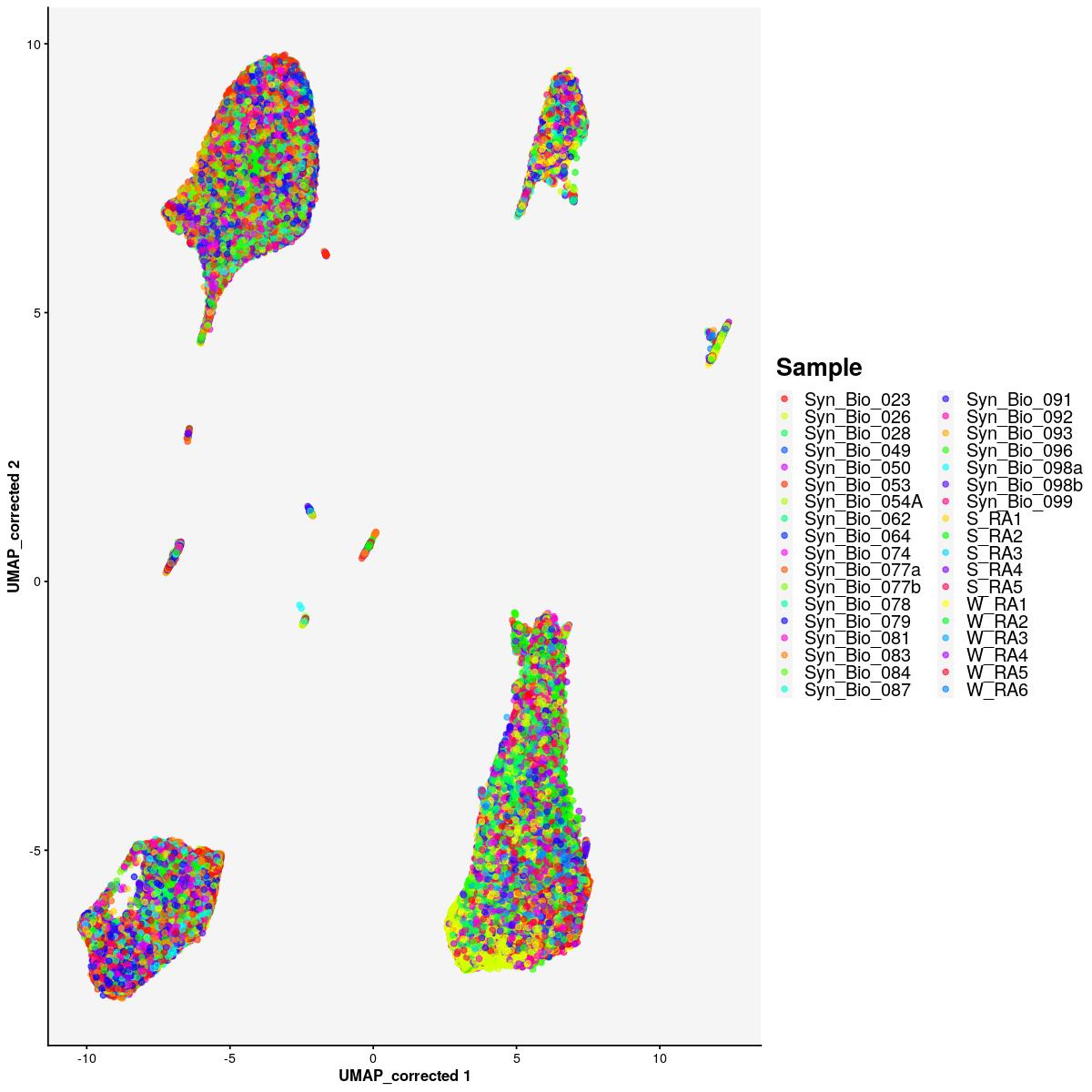
| Version | Author | Date |
|---|---|---|
| 58eeb06 | Reto Gerber | 2023-05-30 |
cat("#### color by Protocol \n\n")color by Protocol
print(fig_ls[['all']]$sfig2c2)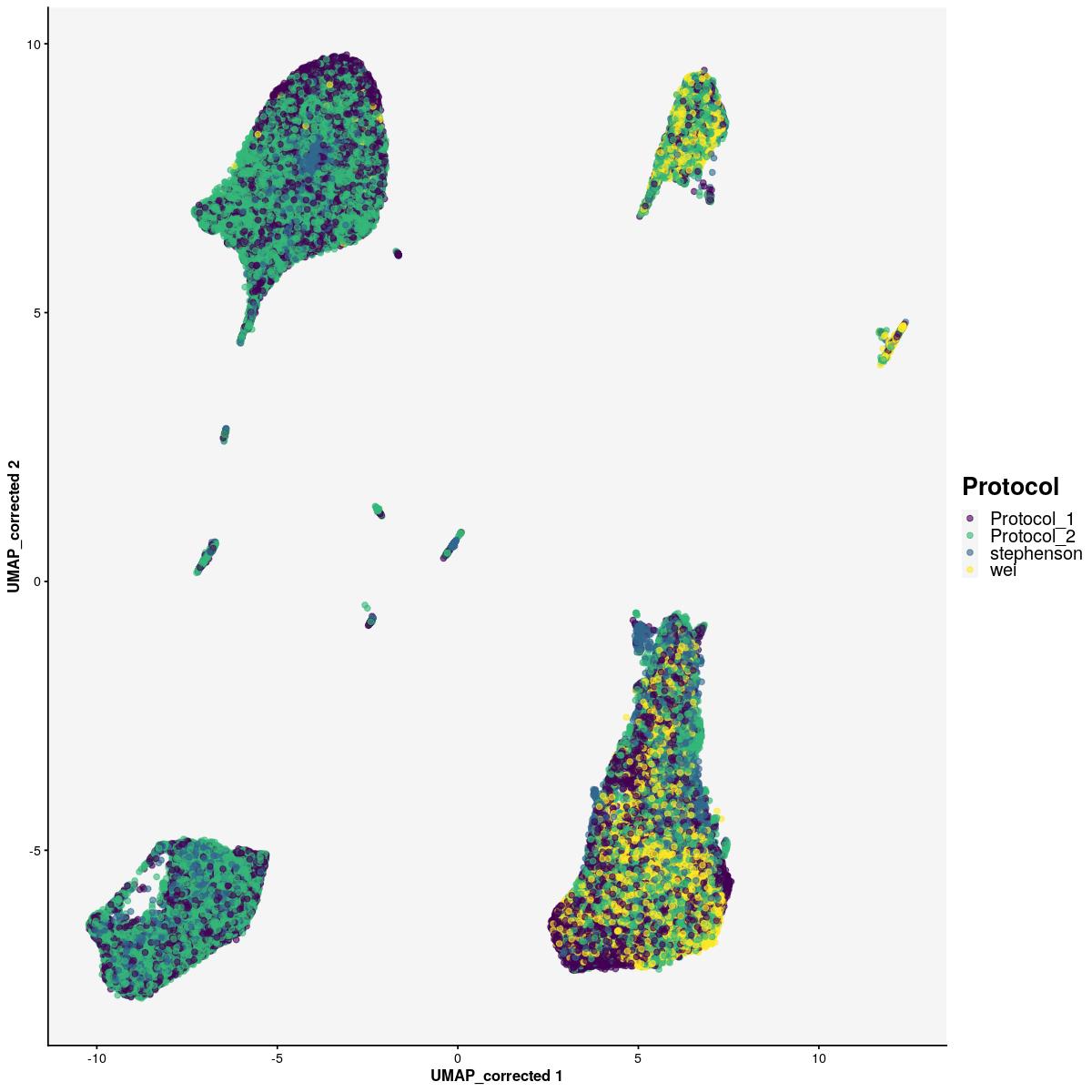
| Version | Author | Date |
|---|---|---|
| 58eeb06 | Reto Gerber | 2023-05-30 |
cat("\n\n### {-}")saveRDS(syn_sce_cms, file = here::here("output",paste0("combined_v",analysis_version,"_sce_hvg_cms.rds")))
# saveRDS(syn_sce_cms, file = here::here("output",paste0("protocol_v",analysis_version,"_sce_batchtest_1.rds")))
# syn_sce_cms <- readRDS(file = here::here("output",paste0("protocol_v",analysis_version,"_sce_batchtest_1.rds")))
sessionInfo()R version 4.0.3 (2020-10-10)
Platform: x86_64-pc-linux-gnu (64-bit)
Running under: Ubuntu 20.04 LTS
Matrix products: default
BLAS/LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.8.so
locale:
[1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C
[3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
[5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=C
[7] LC_PAPER=en_US.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
attached base packages:
[1] parallel stats4 stats graphics grDevices utils datasets
[8] methods base
other attached packages:
[1] gdtools_0.2.3 tidySingleCellExperiment_1.0.0
[3] BiocParallel_1.24.1 bluster_1.0.0
[5] scuttle_1.0.4 igraph_1.2.6
[7] scran_1.18.7 scater_1.18.6
[9] SingleCellExperiment_1.12.0 SummarizedExperiment_1.20.0
[11] Biobase_2.50.0 GenomicRanges_1.42.0
[13] GenomeInfoDb_1.26.7 IRanges_2.24.1
[15] S4Vectors_0.28.1 BiocGenerics_0.36.1
[17] MatrixGenerics_1.2.1 matrixStats_0.58.0
[19] stringr_1.4.0 purrr_0.3.4
[21] ggplot2_3.3.3 dplyr_1.0.4
[23] workflowr_1.6.2
loaded via a namespace (and not attached):
[1] readxl_1.3.1 backports_1.2.1
[3] systemfonts_1.0.1 plyr_1.8.6
[5] lazyeval_0.2.2 kSamples_1.2-9
[7] digest_0.6.27 SuppDists_1.1-9.5
[9] CellMixS_1.6.1 htmltools_0.5.1.1
[11] viridis_0.5.1 fansi_0.4.2
[13] magrittr_2.0.1 openxlsx_4.2.3
[15] limma_3.46.0 svglite_1.2.3.2
[17] colorspace_2.0-0 haven_2.3.1
[19] xfun_0.21 crayon_1.4.1
[21] RCurl_1.98-1.2 jsonlite_1.7.2
[23] glue_1.4.2 gtable_0.3.0
[25] zlibbioc_1.36.0 XVector_0.30.0
[27] DelayedArray_0.16.3 car_3.0-10
[29] BiocSingular_1.6.0 abind_1.4-5
[31] scales_1.1.1 DBI_1.1.1
[33] edgeR_3.32.1 rstatix_0.7.0
[35] Rcpp_1.0.6 viridisLite_0.3.0
[37] dqrng_0.2.1 foreign_0.8-81
[39] rsvd_1.0.3 ResidualMatrix_1.0.0
[41] htmlwidgets_1.5.3 httr_1.4.2
[43] yaImpute_1.0-32 ellipsis_0.3.1
[45] pkgconfig_2.0.3 farver_2.0.3
[47] uwot_0.1.10 locfit_1.5-9.4
[49] here_1.0.1 tidyselect_1.1.0
[51] labeling_0.4.2 rlang_0.4.10
[53] later_1.1.0.1 munsell_0.5.0
[55] cellranger_1.1.0 tools_4.0.3
[57] cli_2.3.0 generics_0.1.0
[59] broom_0.7.4 ggridges_0.5.3
[61] batchelor_1.6.3 evaluate_0.14
[63] yaml_2.2.1 RhpcBLASctl_0.20-137
[65] knitr_1.31 fs_1.5.0
[67] zip_2.1.1 sparseMatrixStats_1.2.1
[69] whisker_0.4 compiler_4.0.3
[71] beeswarm_0.2.3 plotly_4.9.3
[73] curl_4.3 ggsignif_0.6.0
[75] tibble_3.0.6 statmod_1.4.35
[77] stringi_1.5.3 highr_0.8
[79] RSpectra_0.16-0 forcats_0.5.1
[81] lattice_0.20-41 Matrix_1.3-2
[83] vctrs_0.3.6 pillar_1.4.7
[85] lifecycle_1.0.0 RcppAnnoy_0.0.18
[87] BiocNeighbors_1.8.2 data.table_1.13.6
[89] cowplot_1.1.1 bitops_1.0-6
[91] irlba_2.3.3 httpuv_1.5.5
[93] R6_2.5.0 promises_1.2.0.1
[95] gridExtra_2.3 rio_0.5.16
[97] vipor_0.4.5 codetools_0.2-18
[99] assertthat_0.2.1 intrinsicDimension_1.2.0
[101] rprojroot_2.0.2 withr_2.4.1
[103] GenomeInfoDbData_1.2.4 hms_1.0.0
[105] grid_4.0.3 beachmat_2.6.4
[107] tidyr_1.1.2 rmarkdown_2.6
[109] DelayedMatrixStats_1.12.3 carData_3.0-4
[111] git2r_0.28.0 ggpubr_0.4.0
[113] ggbeeswarm_0.6.0